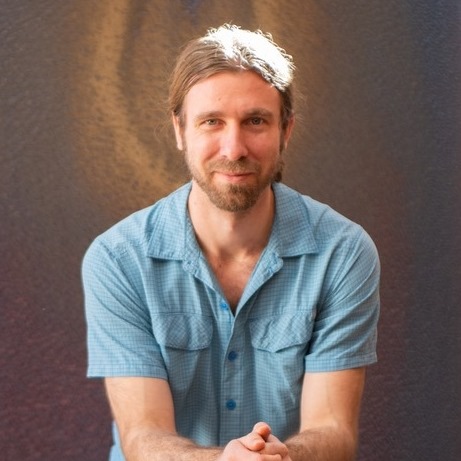
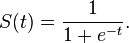Lactose intolerance is the global default. Roughly two-thirds of the world's adults stop producing lactase past childhood, and what counts as a tolerable dose varies hugely between people. The NIH puts the typical threshold at about 12 g of lactose — roughly one cup of milk — without symptoms or with only mild ones1. Below that, most people are fine; above it, dose and individual sensitivity start to matter.
What makes the topic interesting is that the lactose content of dairy products spans nearly five orders of magnitude. A wedge of aged Cheddar has essentially none. A spoonful of Brunost (Norwegian whey cheese) has more than half its mass in lactose. The reason: lactose is water-soluble and lives in the whey, not the curd. Anything that drains whey (cheesemaking, Greek-style yogurt straining) removes lactose. Anything that ferments (aging, live yogurt cultures) converts it to lactic acid. Anything that concentrates whey (Brunost, ricotta, dry milk solids) goes the other way.
The table below is my attempt at a unified reference. All values are grams of lactose per 100 g of product, so they're directly comparable. Each value is footnoted to a single source. Where US (USDA / NIDDK) numbers and non-US databases differ systematically — Feta is the standout — I've gone with the US value.
For mental conversion: 100 g is about 1 cup of milk (which weighs ~244 g, but the lactose-per-100g and per-100mL values are nearly identical for milk), about 3.5 oz of cheese, or roughly 7 tablespoons of butter.
The Safe? column estimates whether a lactose-intolerant adult can comfortably consume a typical US serving (1 cup milk/yogurt, 1 oz cheese, 1 tbsp butter, 2 tbsp cream, ½ cup ice cream, ½ cup cottage cheese / ricotta) with a 3× margin of safety — most people eat more than one serving at a meal, so the rule asks whether 3 servings still fit well under the ~12 g/day NIDDK threshold1:
- Yes — < 1 g per typical serving. Three servings still well under threshold; safe for nearly all lactose-intolerant adults.
- Maybe — 1–4 g per typical serving. One serving is fine for most people; 3 servings approach or hit the threshold.
- No — > 4 g per typical serving. Three servings clearly exceed the threshold; even one serving may cause symptoms in sensitive individuals.
The table
Milks
| Product | Lactose (g/100g) | Safe? | Notes |
|---|---|---|---|
| Whole cow milk (3.25% fat) | 4.812 | No | ≈ 12 g per 244 g cup1 — one cup alone hits threshold |
| Reduced-fat (2%) cow milk | 4.692 | No | |
| Lowfat (1%) cow milk | 4.862 | No | |
| Skim cow milk | 5.052 | No | Removing fat slightly raises lactose-by-mass |
| Buttermilk, lowfat (cultured) | 4.03 | No | Despite the name, US buttermilk is fermented lowfat milk |
| Goat milk | 4.272 | No | |
| Sheep milk | 4.44 | No | |
| Lactose-free milk (Lactaid-style) | <0.14 | Yes | Lactase enzyme has hydrolyzed the lactose into glucose + galactose |
| Evaporated whole milk | ~105 | Maybe | Used in small amounts (2 tbsp ≈ 3 g); a ½ cup recipe portion is "No" |
| Sweetened condensed milk | ~125 | Maybe | A 2 tbsp drizzle is fine; ¼ cup in a recipe is not |
Yogurts and fermented milks
| Product | Lactose (g/100g) | Safe? | Notes |
|---|---|---|---|
| Plain whole-milk yogurt | 4.73 | No | A 6 oz container has ~8 g; live cultures pre-digest some lactose but most remains |
| Plain low-fat yogurt | 4.03 | No | ~6.8 g per 6 oz container |
| Greek yogurt (strained, plain) | ~3.06 | Maybe | Straining removes ~70% of the lactose into the acid whey6 |
| Greek yogurt (nonfat, strained) | ≤0.76 | Yes | |
| Kefir, plain | 4.03 | Maybe | High raw lactose, but live yeast/bacteria reduce symptoms in lactose-intolerant people by 54–71%7 — effective dose is much lower than the raw number suggests |
| Skyr | 2.53 | Maybe | Strained, like Greek yogurt |
| Sour cream | 2.03 | Yes | A 2 tbsp dollop has ~0.6 g |
| Crème fraîche | ~2.44 | Yes | A 2 tbsp serving has ~0.7 g |
Cream and butter
| Product | Lactose (g/100g) | Safe? | Notes |
|---|---|---|---|
| Butter, salted | 0.18 | Yes | An entire stick (113 g) has ~0.1 g; almost all the whey has been churned out |
| Heavy / whipping cream | 2.53 | Yes | A 2 tbsp serving has ~0.75 g; more fat, less aqueous phase, less lactose |
| Half-and-half | ~4.05 | Maybe | A 2 tbsp coffee splash is fine (~1.2 g); 3 splashes approach the threshold |
Fresh and unripened cheeses
| Product | Lactose (g/100g) | Safe? | Notes |
|---|---|---|---|
| Cottage cheese, low-fat | 1.83 | Maybe | A ½ cup serving has ~2 g; 3 servings = 6 g |
| Cream cheese | 3.63 | Maybe | Highest of any common cheese; a 1 oz schmear has ~1 g |
| Ricotta (whey-based) | 2.753 | Maybe | Made from the whey drained off other cheeses; ½ cup has ~3.4 g |
| Mascarpone | 3.03 | Yes | A 1 oz serving has ~0.84 g |
| Chèvre, fresh (goat) | 0.93 | Yes | A 1 oz serving has ~0.25 g |
| Paneer | 0.0023 | Yes | Acid-set; the whey carries off nearly all of it |
| Burrata | ~1.53 | Maybe | A mozzarella shell wrapped around a stracciatella cream filling; lactose is dominated by the filling. ~1 g per typical 1 oz serving |
| Stracciatella (burrata cream filling) | 1.83 | Maybe | Sold as a stand-alone topping; eaten by the spoonful |
| Cheese curds, fresh | 3.03 | Maybe | Wisconsin specialty. Unaged and unpressed, so much of the lactose remains; a small handful (~30 g) has ~1 g |
| Queso fresco | 2.39 | Maybe | Fresh, unaged Mexican cheese. Crumbled on tacos at typical portions (~30 g) gives <1 g |
Soft-ripened and washed-rind cheeses
| Product | Lactose (g/100g) | Safe? | Notes |
|---|---|---|---|
| Brie | <0.00243 | Yes | Mould ripening fully ferments residual lactose |
| Camembert | <0.00243 | Yes | |
| Limburger | <0.00243 | Yes | |
| Taleggio | <0.0013 | Yes |
Pasta filata (stretched-curd) cheeses
| Product | Lactose (g/100g) | Safe? | Notes |
|---|---|---|---|
| Mozzarella, commercial low-moisture | 0.743 | Yes | The kind on US pizza |
| Mozzarella di bufala | 0.353 | Yes | |
| Bocconcini (mini fresh mozzarella) | <0.0013 | Yes | Fresh; lower lactose than commercial mozzarella because there's less moisture to retain whey |
| String cheese | 0.743 | Yes | Mechanically the same product as low-moisture mozzarella; one stick (~28 g) has ~0.2 g |
| Oaxaca (queso Oaxaca) | <0.53 | Yes | Mexican pasta filata cheese; USDA reports ~0 carbohydrates per serving |
| Provolone, dolce or piccante | <0.0013 | Yes |
Hard, semi-hard, and pressed cheeses
These are all functionally lactose-free. The Cheese Scientist database reports values below the analytical detection limit (≈ 1 mg/100 g) for all of them3. Two mechanisms combine: pressing the curds drains nearly all the whey (and with it, the lactose) before aging even begins; for cheeses aged >2 months, residual lactose is further fermented to lactic acid, as confirmed in the Stelios et al. PDO survey10. Even young pressed cheeses like Havarti and Pepper Jack therefore come in at trace levels.
My current and previous choices for the tools I use.
- Android Apps
- Workout tracker: FitBod (but probably not for much longer - it doesn't have a data export option)
- Bathroom
- Toothbrush: Philips One by Sonicare - compact, lightweight, good battery, charges with a USB-A to USB-C cable. The only improvement here would be the ability to charge with a real USB-C cable.
- Fingernail clipper: Seki Edge
- Nail file: 3 Swords Germany
- Electronics
- Large power adapter: Anker Prime 67W USB C Charger
- Small power adapter: Anker Nano Charger
- Wireless charger: TOZO W1 Wireless Charger 15W Max
- USB-C cable: Anker
- Wireless car charger: APPS2Car
- Car USB-C charger: Roadress
- USB-C magnetic breakaways: TiMOVO
- Keyboard: nuphy Air75
- USB-C to USB-A adapter: Syntech
- Computer camera cover: CloudValley
- Hiking
- Climbing
- Belay glasses: BG Climbing
- PPE
- Games
- Misc
- Micro-knife - has passed through TSA many times unmolested
- Kitchen
An answer to this question on Stack Overflow.
Question
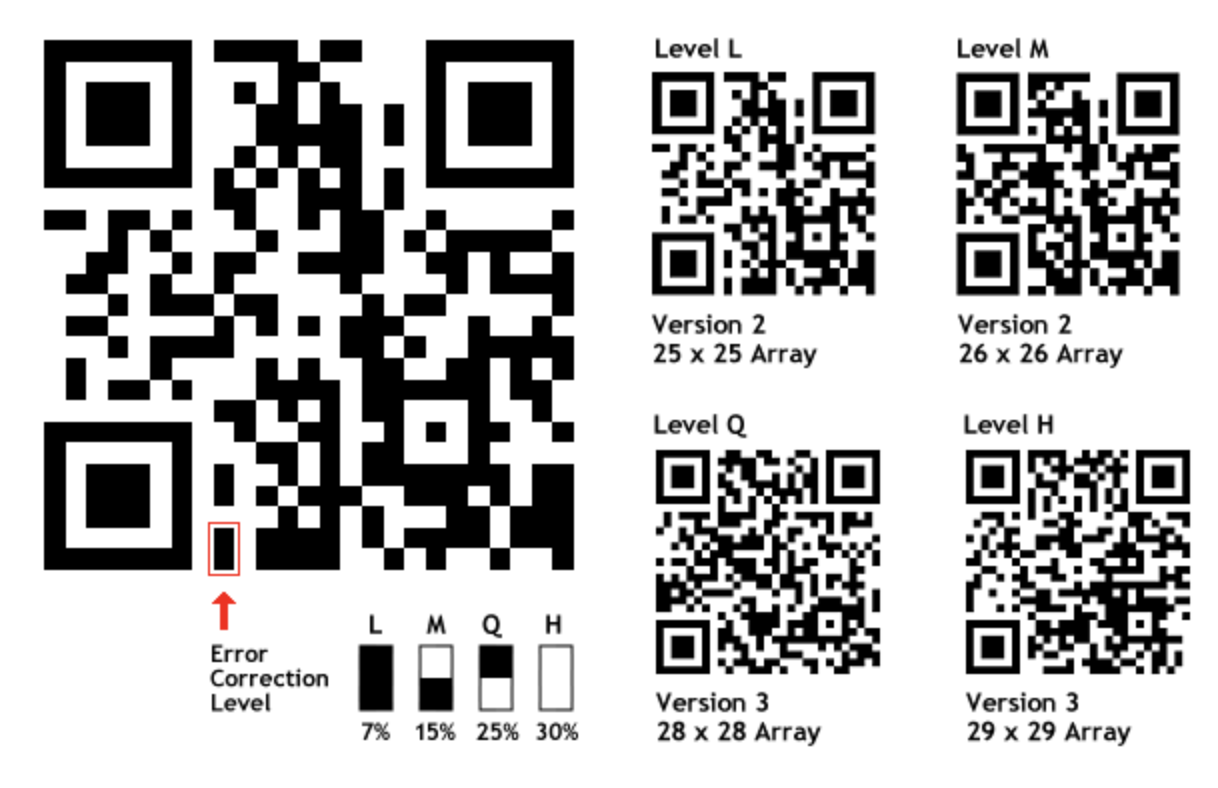
I'm not a specialist, but as far as I know, a bit of information in a QR-code is coded more than once, and it is defined as the redundancy level
How can I estimate a QR-code redundancy level ? Is where an mobile app or a website where I can test my QR-code redundancy level easily ? If not, is it an easy algorithm that I can implement ?
Redundancy is sorted in different categories according to this website, but I'd like to have the direct percentage value if possible
Answer
QR codes contain a couple of bits which indicate the error correction level, as depicted below (source):
An answer to this question on Stack Overflow.
Question
The sigmoid function is defined as
S(t) = 1 / (1 + e^(-t))
(where ^ is pow)
I found that using the C built-in function exp() to calculate the value of f(x) is slow. Is there any faster algorithm to calculate the value of f(x)?
Answer
This question has significantly more detail including these benchmarked results:
name rms_error maxdiff time_us speedup samples
logistic_with_tanh 5.9496e-02 1.5014e-01 0.0393 0.5076 200000001
logistic_with_atan 3.9051e-02 9.6934e-02 0.0321 0.6211 200000001
logistic_with_erf 6.5068e-02 1.6581e-01 0.0299 0.6676 200000001
logistic_fexp_quintic_approx 1.2921e-07 5.9050e-07 0.0246 0.8118 200000001
logistic_product_approx_float128 8.7032e-04 1.7217e-03 0.0209 0.9523 200000001
logistic_with_exp_no_overflow 4.7660e-17 1.6653e-16 0.0198 1.0084 200000001
logistic_product_approx128 8.7032e-04 1.7211e-03 0.0164 1.2187 200000001
log_w_approx_exp_no_overflow128 8.7193e-04 1.7211e-03 0.0158 1.2640 200000001
logistic_with_sqrt 8.3414e-02 1.1086e-01 0.0146 1.3662 200000001
log_w_approx_exp_no_overflow16 6.9726e-03 1.4074e-02 0.0141 1.4114 200000001
log_w_approx_exp_no_overflow16_clamped 6.9726e-03 1.4074e-02 0.0141 1.4153 200000001
logistic_schraudolph_approx 1.5661e-03 8.9906e-03 0.0138 1.4497 200000001
logistic_with_abs 6.0968e-02 8.2289e-02 0.0134 1.4936 200000001
logistic_orig 0.0000e+00 0.0000e+00 0.0199 ------ 200000001
An answer to this question on Stack Overflow.
Question
Problem background
I have a 2D array
Map[Height][Width]that stores aboolto represent each cell.There exists two non-overlapping regions
RegionAandRegionB.Each region is a list of unique adjacent integer co-ordinates.
The number of co-ordinates in
RegionAandRegionBaremandnrespectivelyA co-ordinate in
RegionAwill never be horizontally or vertically adjacent to a coordinate inRegionB(i.e. the regions do not touch)The bounding box of
RegionAcould overlap with the bounding box ofRegionBRegionAandRegionBmay have holesRegionAmay or may not surroundRegionB(and vise versa)if a co-ordinate (X, Y) is part of a region then
Map[Y][X] == 1, otherwise its zero.
Problem
I'm looking for an algorithm to determine the two co-ordinates A and B that have the minimum Euclidean distance, where A is a member of RegionA and B is a member of RegionB.
The brute force method has a time complexity of O(mn).
Is there a more time-efficient algorithm (preferably better than O(mn)) for solving this problem?
The name + link, code, or description of one instance of such an algorithm would be greatly appreciated and will be accepted as an answer.
C++ code to create two random regions A and B
The region generation code is immaterial to the problem being asked. and I am only interested in how close the regions are to each other. In the project I am working on I used cellular automata to create the regions. I can include the code for this if necessary. The Region construction procedure exists only to create relevant examples.
#include <cstdlib>
#include <cstring>
#include <vector>
#include <cstdio>
#include <map>
#include <ctime>
#include <cmath>
constexpr int Width = 150;
constexpr int Height = 37;
bool Map[Height][Width];
struct coord {
int X, Y;
coord(int X, int Y): X(X), Y(Y) {}
bool operator==(const coord& Other) {
return Other.X == X && Other.Y == Y;
}
bool IsAdjacentTo(const coord& Other){
int ManhattanDistance = abs(X - Other.X) + abs(Y - Other.Y);
return ManhattanDistance == 1;
}
bool IsOnMapEdge(){
return X == 0 || Y == 0 || X == Width - 1 || Y == Height - 1;
}
};
using region = std::vector<coord>;
void ClearMap(){
std::memset(Map, 0, sizeof Map);
}
bool& GetMapBool(coord Coord) {
return Map[Coord.Y][Coord.X];
}
int RandInt(int LowerBound, int UpperBound){
return std::rand() % (UpperBound - LowerBound + 1) + LowerBound;
}
region CreateRegion(){
std::puts("Creating Region");
ClearMap();
region Region;
int RegionSize = RandInt(50, Width * Height / 2);
Region.reserve(RegionSize);
while (true){
Region.emplace_back(
RandInt(1, Width - 1),
RandInt(1, Height - 1)
);
if (!Region[0].IsOnMapEdge())
break;
else
Region.pop_back();
}
GetMapBool(Region[0]) = 1;
while (Region.size() != RegionSize){
coord Member = Region[RandInt(0, Region.size() - 1)];
coord NeighbourToMakeAMember {
RandInt(-1, 1) + Member.X,
RandInt(-1, 1) + Member.Y
};
if (!Member.IsOnMapEdge() && !GetMapBool(NeighbourToMakeAMember) && Member.IsAdjacentTo(NeighbourToMakeAMember)){
GetMapBool(NeighbourToMakeAMember) = 1;
Region.push_back(NeighbourToMakeAMember);
};
}
std::puts("Created Region");
return Region;
}
std::pair<region, region> CreateTwoRegions(){
bool RegionsOverlap;
std::pair<region, region> Regions;
do {
Regions.first = CreateRegion();
Regions.second = CreateRegion();
ClearMap();
for (coord Member : Regions.first){
GetMapBool(Member) = 1;
}
RegionsOverlap = 0;
for (coord Member : Regions.second){
if (GetMapBool(Member)){
ClearMap();
std::puts("Regions Overlap/Touch");
RegionsOverlap = 1;
break;
} else {
GetMapBool(Member) = 1;
}
}
} while (RegionsOverlap);
return Regions;
}
void DisplayMap(){
for (int Y = 0; Y < Height; ++Y){
for (int X = 0; X < Width; ++X)
std::printf("%c", (Map[Y][X] ? '1' : '-'));
std::puts("");
}
}
int main(){
int Seed = time(NULL);
std::srand(Seed);
ClearMap();
auto[RegionA, RegionB] = CreateTwoRegions();
DisplayMap();
}
Problem illustration
What are the co-ordinates of the points A and B that form the minimum Euclidean distance between the two regions?
Answer
Notes on finding the nearest points between two regions
As others have mentioned, you can improve on finding the closest points between the two regions of n and m points by first filtering each region's points to only those which border unassigned cells and then doing an all-pairs search between the boundaries. For a roughly rectangular region this would take O(m + n + 16 √m √n) time: two linear filters and then the all-pairs search. I've implemented an example of this search below.
But this will not save you any time if your regions are shaped like this:
########
# #
# #### #
# # # #
# # ## #
# # #
# ######
If you are using larger datasets and can tolerate preprocessing you can get further acceleration using a 2D k-d tree.
Construction takes O(n log n) time after which you can search the tree in O(log n) time for m queries. So the total time to find the closest points is then O(m log n + n log n) or O(n log m + m log m).
You can do some benchmarking to determine if it is better to build the k-d tree for the larger or the smaller of the regions. Remember that this answer may change depending on how many times you're able to reuse the same k-d tree for queries against different regions.
For small datasets you'll probably find that the all-pairs comparison is faster because it has much greater cache locality.
Note that implementing a k-d tree is tricky, so you'll probably want to use an existing library or definitely use test cases if you're rolling your own.
Another algorithm
If we happen to know that the regions are within a distance d of each other we can get an even faster algorithm.
For one of the regions we can build a hash table where a reference to each member cell is stored at a hash location (c.y // d) * height + (c.x // d). To find the nearest neighbor to a query point we then do an all-pairs check against the points referenced in the associated "meta-cell" and as well as the meta-cell's neighbors.
Notes on region building
On the off-chance you're trying to grow the regions quickly, but their boundaries can overlap, this should do the trick.
Note that your question can be improved by stating why you are trying to calculate a thing. If you want to know the closest-cell so you have a primitive to build regions more quickly, then reconsidering how region building works might be a better use of your time, and it's what I address here.
If region generation is immaterial and you're only interested in how close they are to each other, then focus on defining, conceptually, what a region is and be very clear that your region construction procedure exists only to create relevant examples.
The fundamental trick we'll use to accelerate region construction is to make a space-time tradeoff. Rather than storing the map as a set of boolean values, we'll store it as a set of 8-bit integers. We reserve a few of these integers for special purposes:
- 0 - An unassigned space that a region may be built on
- 1 - A buffer cell that isn't part of any region, but close to one. This is not available for building on.
- 2, 3, 4, ... - valid region labels.
Thus, the algorithm for building a region becomes simple:
- Find a valid seed cell
- Grow outward from the seed cell via 4-connected cells until the desired number of cells is reached
- A new Region Cell can only be placed in an Unassigned Space
- Iterate over Region Cells marking their 8-connected neighbors as Buffer cells.
- Note that, if we add the new Buffer cells to the list of Region Cells and repeat the last step, we can grow arbitrarily large buffers between regions.
A few other parts of your code needed cleaning. Types are typically defined using Camel Case and variables use snake case. I prefer snake_case for functions as well. See a style guide for more opinions.
Do not use global variables they are bad and, in the fullness of time, they will bring you nothing but sorrow.
You're using C++, avoid using headers such as stdio, stdlib, and cstring. They provide the C way of doing things, which often come with disadvantageous such as diminished type safety.
Any time your if statements have a lot of conditions, ask yourself whether they can be simplified by inverting one or more of the conditions (that is, switching is_good() to !is_good()) and doing an early exit. This can make your code much easier to read and reasonable about.